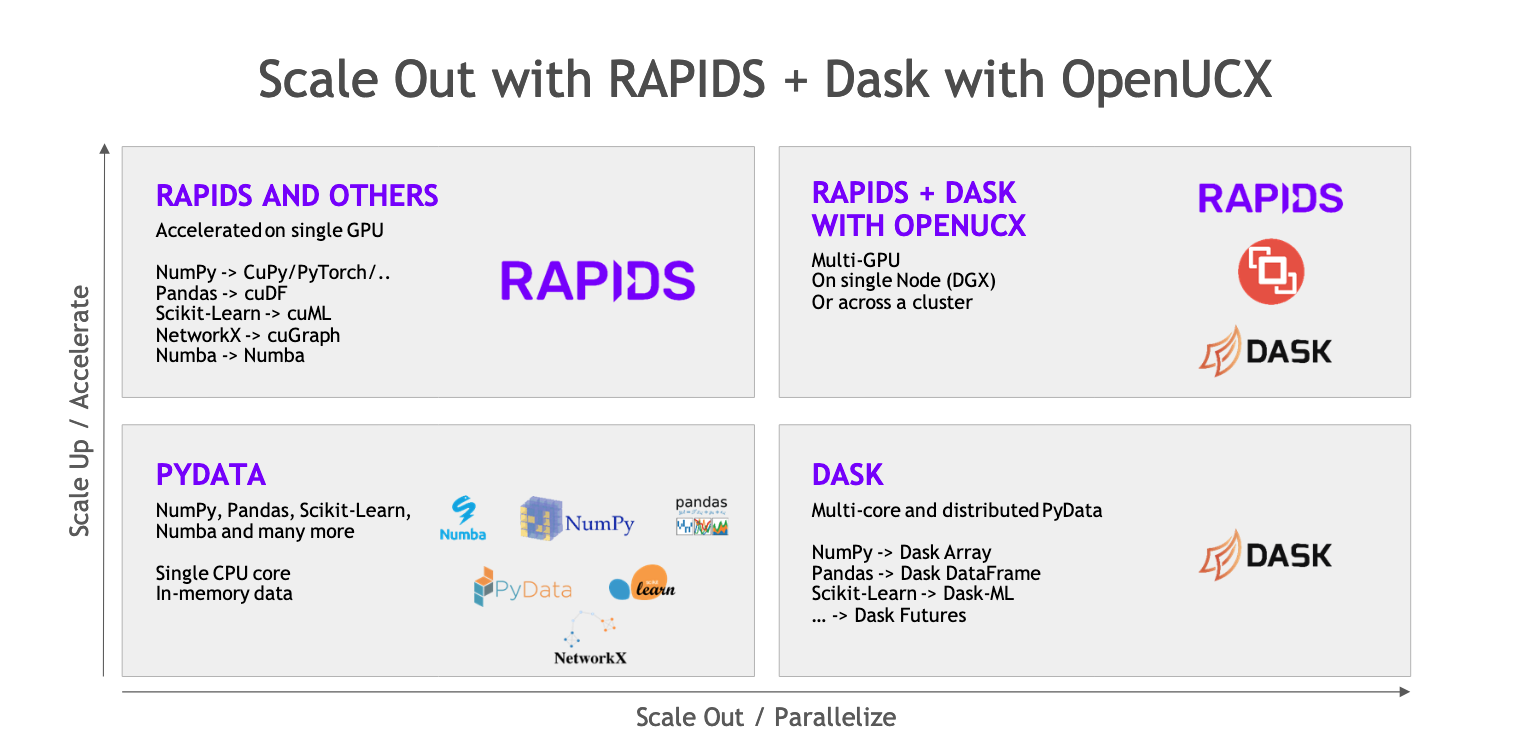
Both Dask and RAPIDS are Python libraries to scale your workflow and empower you to process more data and leverage more compute resources. Both use interfaces modeled after the PyData ecosystem, making them familiar to most data practitioners.
So what’s the difference?
Dask allows you to scale out your workload onto multiple processors and multiple machines by creating an abstract representation of your code and then distributing that onto a cluster.
RAPIDS allows you to scale up your workload by reimplementing low-level parts of common open source APIs to run on NVIDIA GPUs giving you faster execution.
Together, Dask and RAPIDS allow you to scale up and scale out production Python workloads at the same time.
Dask
Dask enables you to distribute your work by creating a Directed Acyclic Graph (DAG) representation of your code at runtime. This graph is then passed to a scheduler that allocates individual tasks to worker processes. These worker processes can be on a single machine, allowing you to make use of all your CPU cores, or on many machines allow you to scale to hundreds or even thousands of CPU cores.
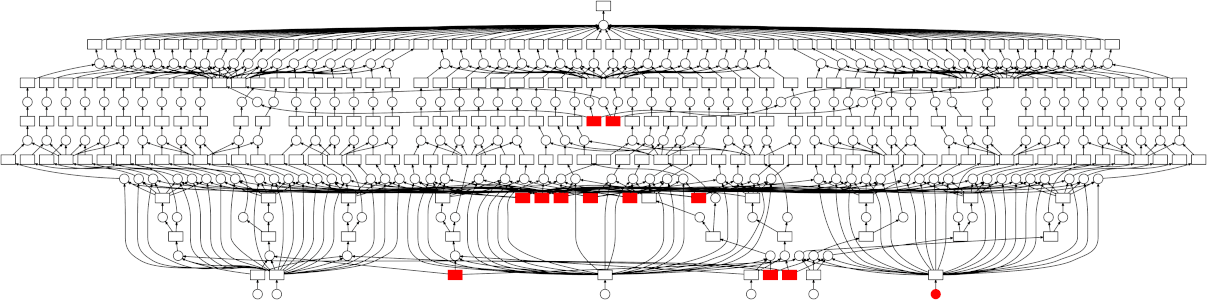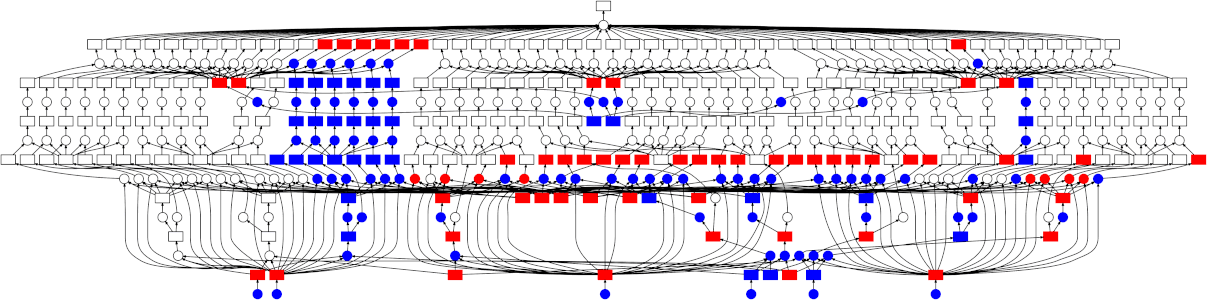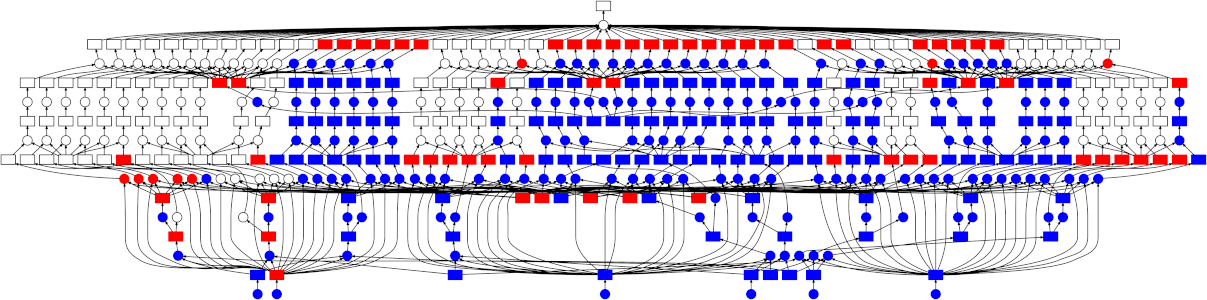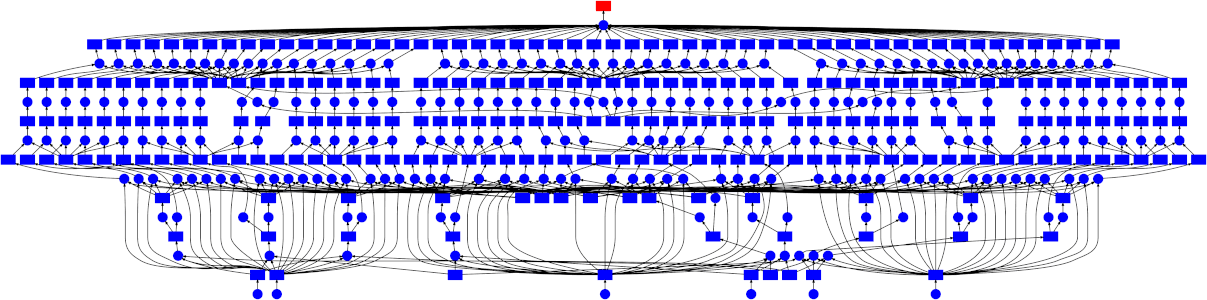
While Dask can be used directly to create these graphs using the Delayed interface there are also implementations of common Python APIs which have already been converted over.
Take pandas for example. pandas is a popular library for working with Dataframes in Python. However, it is single-threaded and the Dataframes you are working on must fit within memory.
import pandas as pd
df = pd.read_csv("/path/to/my/data-*.csv") # Your CSV data must be smaller than your memory
Dask has a sub-package called dask.dataframe which follows most of the same API as pandas but instead breaks your Dataframe down into partitions that can be operated on in parallel and can be swapped in and out of memory. Dask uses pandas under the hood, so each partition is a valid pandas Dataframe.
import dask.dataframe as dd
ddf = dd.read_csv("/path/to/my/data-*.csv") # Your CSV files can be larger than memory
The overall Dask Dataframe can then scale-out and use multiple cores or multiple machines.
RAPIDS
RAPIDS is a collection of GPU-accelerated open source libraries that follow the APIs of other popular open source packages.
To continue with our pandas theme, RAPIDS has a package called cuDF, which mirrors the same API as pandas. However, cuDF stores Dataframes in GPU memory and uses the GPU to perform computations.
import cudf
gdf = cudf.read_csv("/path/to/my/data-*.csv") # Your CSV data must be smaller than your GPU memory
As GPUs can accelerate computations this can lead to performance benefits for your Dataframe operations and enables you to scale up your workflow.
RAPIDS and Dask
RAPIDS and Dask are integrated, so Dask is considered a component of RAPIDS. Many of NVIDIA’s RAPIDS developers are also Dask developers. So instead of having a Dask Dataframe made up of individual pandas Dataframes you could instead have one made up of cuDF Dataframes to run on GPUs. This is possible because they follow the same pandas-like API.
import dask_cudf
gdf = dask_cudf.read_csv("/path/to/my/data-*.csv") # Your CSV files can be larger than GPU memory and you can use multiple GPUs to process this data
This way you can both scale up by using a GPU and also scale-out using multiple GPUs on multiple machines.
Recap
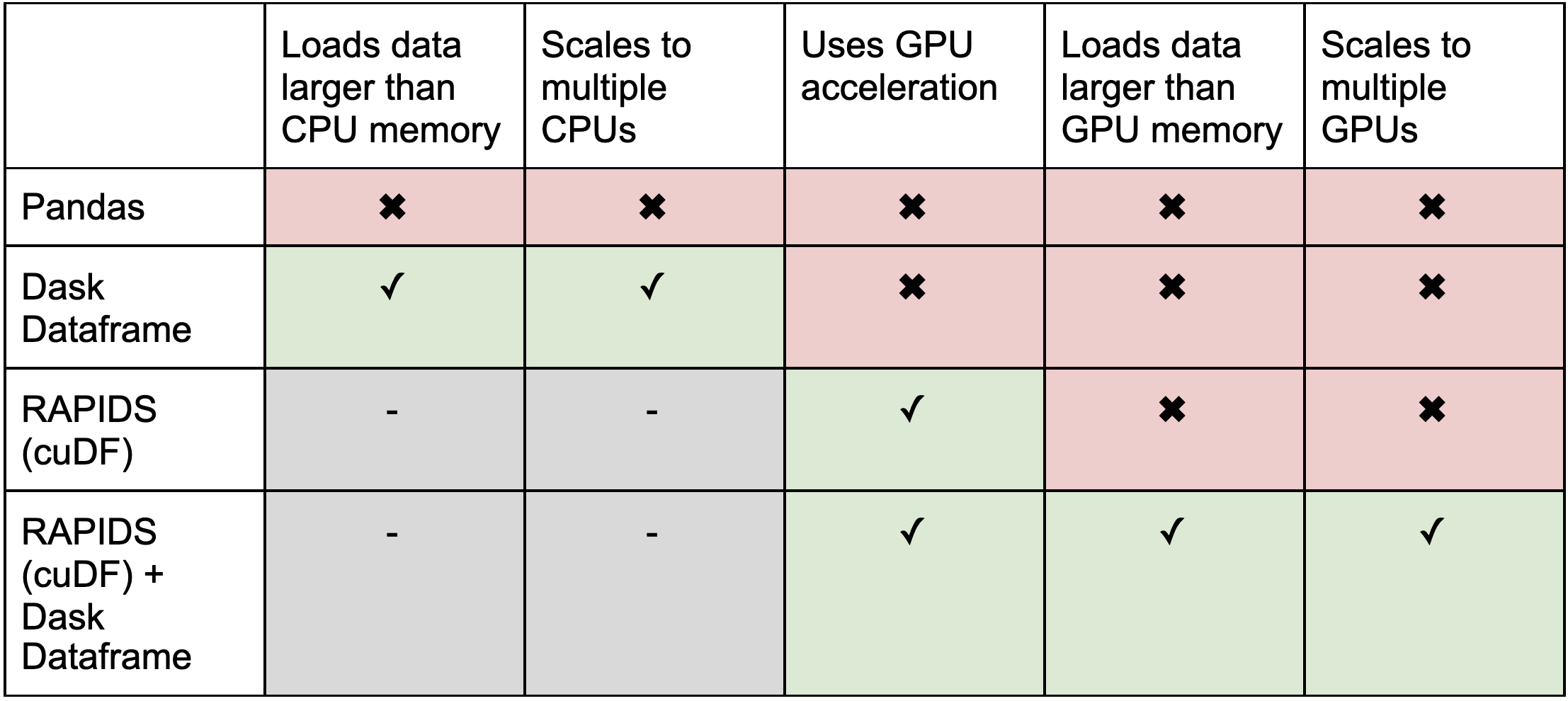

Further Reading
In this post, we’ve only scratched the surface of what RAPIDS and Dask can do. They both also support NumPy style array workflows with dask.array and CuPy and scikit-learn machine learning workflows with dask-ml and cuML.
 Jacob Tomlinson
Jacob Tomlinson